
HOMER
Software for motif discovery and next-gen sequencing analysis
Finding Enriched Motifs in Genomic Regions (findMotifsGenome.pl)
HOMER was initially developed to automate the process of
finding
enriched motifs in ChIP-Seq peaks. More generally,
HOMER analyzes
genomic positions, not limited to only ChIP-Seq peaks, for
enriched
motifs. The main idea is that all the user really
needs is a file
containing genomic coordinates (i.e. a HOMER peak file
or BED
file),
and
HOMER
will
generally
take
care
of
the
rest.
To
analyze a peak file for
motifs, run the following command:findMotifsGenome.pl
<peak/BED
file>
<genome>
<output
directory> -size #
[options]
i.e. findMotifsGenome.pl ERpeaks.txt hg18 ER_MotifOutput/ -size 200 -mask
i.e. findMotifsGenome.pl ERpeaks.txt hg18 ER_MotifOutput/ -size 200 -mask
A variety of output files will be placed in the <output directory>, including html pages showing the results. The "-mask" is optional and tells the program to use the repeat-masked sequence. (The old shorthand hg18r will also work). The -size parameter is now mandatory when running findMotifsGenome.pl to avoid confusion - plus it's always a good idea to know exactly what size the regions you are analyzing are.
The findMotifsGenome.pl program is a wrapper that helps set up the data for analysis using the HOMER motif discovery algorithm. By default this will perform de novo motif discovery as well as check the enrichment of known motifs. If you have not done so already, please look over this page describing how HOMER analyzes sequences for enriched motifs.
An important prerequisite for analyzing genomic motifs is that the appropriate genome must by configured for use with HOMER. In version v3.1, HOMER now handles custom/arbitrary genomes. Instead of intalling/configuring a genome, you can specify the path to a file or directory containing the genomic sequence in FASTA format. The genome can be in a single FASTA file, or you specify a directory where where each chromosome can be in a separate file (named chrXXX.fa or chrXXX.fa.masked). In either case, the FASTA headers must contain the chromosome names followed by white space, i.e. ">chr blahblahblah", not ">chr1-blahblahblah", or prefereably only ">chr1". (also note that homer will create a "preparsed/" directory where the genome is, so make sure you have write permissions in the genomic directory.
Selecting the size of the region for motif finding
(-size # or -size given, default: 200)
This is one of the most important parameters and also a source of confusion for many. If you wish to find motifs using your peaks using their exact sizes, use the option "-size given"). However, for Transcription Factor peaks, most of the motifs are found +/- 50-75 bp from the peak center, making it better to use a fixed size rather than depend on your peak size.
Acceptable Input files
findMotifsGenome.pl
accepts HOMER
peak files
or BED files:
HOMER peak files should have at minimum 5 columns (separated by TABs, additional columns will be ignored):
Mac Users: If using a EXCEL to prepare input files, make sure to save files as a "Text (Windows)" if running MacOS - saving as "Tab delimited text" in Mac produces problems for the software. Otherwise, you can run the script "changeNewLine.pl <filename>" to convert the Mac-formatted text file to a Windows/Dos/Unix formatted text file.
If errors occur, it is likely that the file is not in the correct format, or the first column is not actually populated with unique identifiers.
HOMER peak files should have at minimum 5 columns (separated by TABs, additional columns will be ignored):
- Column1: Unique Peak ID
- Column2: chromosome
- Column3: starting position
- Column4: ending position
- Column5: Strand (+/- or 0/1, where 0="+", 1="-")
- Column1: chromosome
- Column2: starting position
- Column3: ending position
- Column4: Unique Peak ID
- Column5: not used
- Column6: Strand (+/- or 0/1, where 0="+", 1="-")
Mac Users: If using a EXCEL to prepare input files, make sure to save files as a "Text (Windows)" if running MacOS - saving as "Tab delimited text" in Mac produces problems for the software. Otherwise, you can run the script "changeNewLine.pl <filename>" to convert the Mac-formatted text file to a Windows/Dos/Unix formatted text file.
If errors occur, it is likely that the file is not in the correct format, or the first column is not actually populated with unique identifiers.
Custom Background Regions
Since HOMER uses a
differential
motif discovery algorithm, different types of background
sequences can
be chosen to produce different results. For example,
you may want
to compare the ChIP-Seq peaks specific in one cell type
versus the
peaks that are specific to another. To do this,
create a second
peak/BED file and use it with the argument "-bg <peak/BED file>".
HOMER
will still try to normalize the background to remove
GC-bias and
will also perform autonormalization (see below). You
can turn off
the normalization with ("-noweight"
and/or
"-nlen 0").
How findMotifsGenome.pl works
There are a series of steps
that
the program goes through to find quality motifs:
1. Verify peak/BED file
2. Extract sequences from the genome corresponding to the regions in the input file, filtering sequences that are >70% "N"
3. Calculate GC/CpG content of peak sequences.
4. Preparse the genomic sequences of the selected size to serve as background sequences.
5. Randomly select background regions for motif discovery.
6. Autonormalization of sequence bias.
7. Check enrichment of known motifs
8. de novo motif finding
1. Verify peak/BED file
HOMER makes sure you have
valid
peaks, and checks to make sure you have unique peak
identifiers.
If there are replicates, it will inform you, and will
add numbers to
peak names to ensure they are unique for downstream
analysis.
2. Extract sequences from the genome corresponding to the regions in the input file, filtering sequences that are >70% "N"
This step is pretty self
explanatory. If you wish to extract sequences from
a genome for
any reason, check out homerTools. HOMER
will also trash
sequences that are predominately "N". If you feel
you are
throwing away too many sequences, try running findMotifsGenome.pl
on an unmasked
genome.
3. Calculate GC/CpG content of peak sequences.
CpG Islands are the single
biggest source of sequence content bias in mammalian
genomes, and are
unfortunately found near transcription start sites,
where all the
action is! By default, HOMER tracks GC% (use "-cpg" to use CpG%).
4. Preparse the genomic sequences of the selected size to serve as background sequences.
This step is only done the
first
time you find motifs from regions of a given size ("-size <#>").
HOMER takes
regions near the TSS of genes (+/- 50kb) and splits them
into regions
of the indicated size. It then calculates their
GC/CpG% and
stores them for later use to speed up execution the next
time you
search for motifs from similar sized regions.
5. Randomly select background regions for motif discovery.
Since HOMER is a
differential
motif discovery algorithm, it must use background
sequence regions as a
control. By default, HOMER selects enough random
background
regions such that the total number of regions is 50000
or 2x the total
number of peaks, which ever is larger (to change use "-N <#>").
The more total
sequence that is used, the slower the program will run,
but you want to
make sure there is enough background regions to reliably
estimate motif
frequency. HOMER attempts to select background
regions that match
the GC-content distribution of the input sequences (in
5%
increments). For example, if your input regions
are extremely
GC-rich, HOMER will select random regions from GC-rich
regions of the
genome as a control.
If custom background regions are provided ("-bg <peak/BED file>"), HOMER will automatically ensure that these regions do NOT overlap with the target regions (using mergePeaks). Custom regions will still be normalized for GC-content.
If custom background regions are provided ("-bg <peak/BED file>"), HOMER will automatically ensure that these regions do NOT overlap with the target regions (using mergePeaks). Custom regions will still be normalized for GC-content.
6. Autonormalization of sequence bias.
Autonormalization is a
unique
procedure provided by HOMER that attempts to remove bias
introduced by
lower-order oligo sequences. It works by assuming
your targets
regions and background regions should not have an
imbalance in 1-mers,
2-mers, 3-mers, etc. The maximum length of oligo
that is
autonormalized is specified by "-nlen
<#>" (default is 3, to disable use "-nlen 0"). For
example, there
should not be significantly more A's in the target
sequences that in
the background. After calculating the imbalances
for each oligo,
it adjusts the weights of each background sequence by a
small amount to
help normalize any imbalance. If target sequences
are rich in A,
then background sequences that contain many A's will be
assigned higher
weights while those with very few A's will be assigned
lower
weights. The weights are incremented by only small
amounts and
the procedure repeated many times in a hill climbing
optimization. This procedure helps remove some of
the sequence
bias associated with certain genomic regions, or bias
that may have
been introduced by biased experimental results such as
biased
sequencing.
7. Check enrichment of known motifs
HOMER screens it's library
of
reliable motifs against the target and background
sequences for
enrichment, returning motifs enriched with a p-value
less than
0.05.
The known motif enrichment is performed first since it
is usually
faster, and gives a faster look at what's enriched in
your target
regions. Know motif enrichment will be reported to
the
"knownResults.html" file in the output directory.
8. de novo motif finding
Best saved for last.
By
default, HOMER will search for motifs of len 8, 10, and
12 bp (change
using -len
<#,#,#> with
no spaces between the numbers, i.e. "-len
6,10,15,20"). For a more detail
description of the motif
discovery algorithm, see here. Output from the de
novo motif
finding will be displayed in the "homerResults.html"
file.
findMotifsGenome.pl Output
A full description of motif
finding output and the output can be found here.
Several files are produced in the output directory:
Several files are produced in the output directory:
homerMotifs.motifs<#>
:
these are the output files from the de novo motif
finding, separated by
motif length, and represent separate runs of the
algorithm.
homerMotifs.all.motifs : Simply the concatenated file composed of all the homerMotifs.motifs<#> files.
motifFindingParameters.txt : command used to execute findMotifsGenome.pl
knownResults.txt : text file containing statistics about known motif enrichment (open in EXCEL).
seq.autonorm.tsv : autonormalization statistics for lower-order oligo normalization.
homerResults.html : formatted output of de novo motif finding.
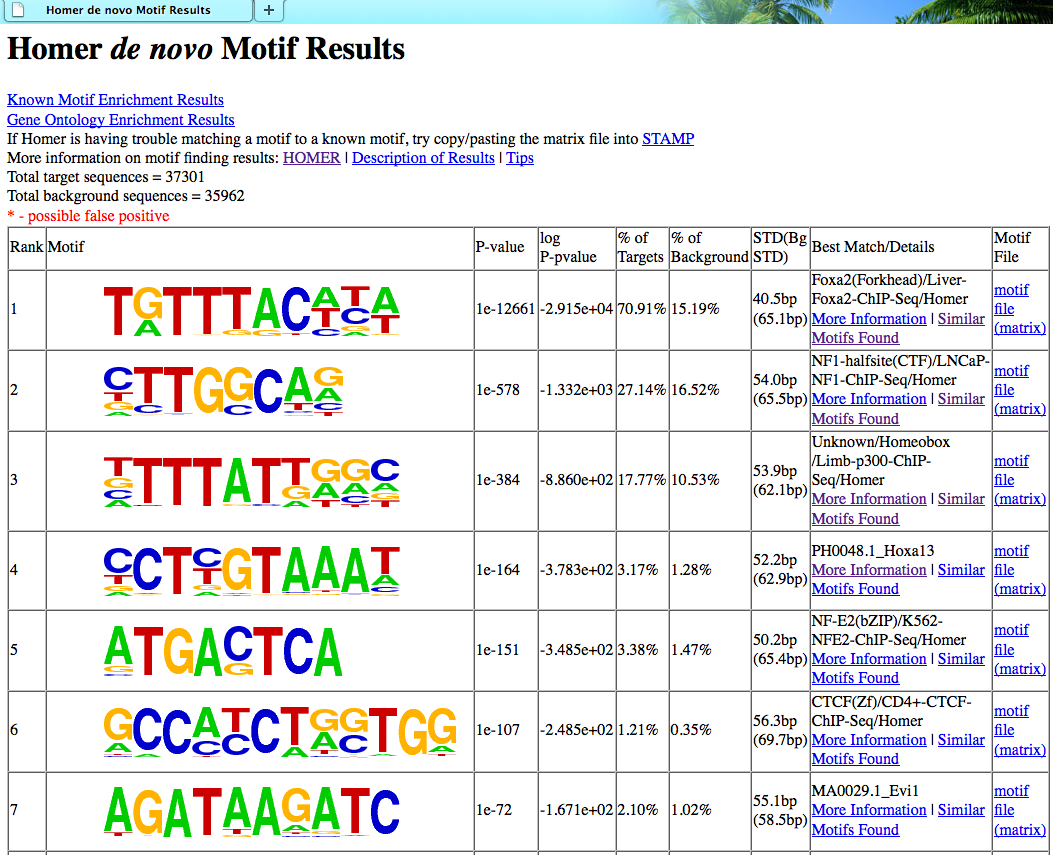
homerResults/ directory: contains files for the homerResults.html webpage, including motif<#>.motif files for use in finding specific instance of each motif.
knownResults.html : formatted output of known motif finding.
knownResults/ directory: contains files for the knownResults.html webpage, including known<#>.motif files for use in finding specific instance of each motif.
homerMotifs.all.motifs : Simply the concatenated file composed of all the homerMotifs.motifs<#> files.
motifFindingParameters.txt : command used to execute findMotifsGenome.pl
knownResults.txt : text file containing statistics about known motif enrichment (open in EXCEL).
seq.autonorm.tsv : autonormalization statistics for lower-order oligo normalization.
homerResults.html : formatted output of de novo motif finding.

homerResults/ directory: contains files for the homerResults.html webpage, including motif<#>.motif files for use in finding specific instance of each motif.
knownResults.html : formatted output of known motif finding.
knownResults/ directory: contains files for the knownResults.html webpage, including known<#>.motif files for use in finding specific instance of each motif.
Interpreting motif finding results
The format of the output
files
generated by findMotifsGenome.pl
are identical to those generated by the promoter-based
version findMotifs.pl
(description).
In general, when analyzing ChIP-Seq / ChIP-Chip peaks you should expect to see strong enrichment for a motif resembling the site recognized by the DNA binding domain of the factor you are studying. Enrichment p-values reported by HOMER should be very very significant (i.e. << 1e-50). If this is not the case, there is a strong possibility that the experiment may have failed in one way or another. For example, the peaks could be of low quality because the factor is not expressed very high.
Practical Tips for Motif finding
In general, when analyzing ChIP-Seq / ChIP-Chip peaks you should expect to see strong enrichment for a motif resembling the site recognized by the DNA binding domain of the factor you are studying. Enrichment p-values reported by HOMER should be very very significant (i.e. << 1e-50). If this is not the case, there is a strong possibility that the experiment may have failed in one way or another. For example, the peaks could be of low quality because the factor is not expressed very high.
Practical Tips for Motif finding
Important motif finding parameters
Masked vs. Unmasked Genome
("-mask" or hg18 vs. hg18r)
Region Size ("-size <#>", "-size <#>,<#>", "-size given", default: 200)
Motif length ("-len <#>" or "-len <#>,<#>,...", default 8,10,12)
Mismatches allowed in global optimization phase ("-mis <#>", default: 2)
Number of CPUs to use ("-p <#>", default 1)
Number of motifs to find ("-S <#>", default 25)
Normalize CpG% content instead of GC% content ("-cpg")
Region level autonormalization ("-nlen <#>", default 3, "-nlen 0" to disable)
Motif level autonormalization (-olen <#>, default 0 i.e. disabled)
User defined background regions ("-bg <peak file of background regions>")
Hypergeometric enrichment scoring ("-h")
Find enrichment of individual oligos ("-oligo").
Force findMotifsGenome.pl to re-preparse genome for the given region size ("-preparse").
Only search for motifs on + strand ("-norevopp")
Search for RNA motifs ("-rna")
Mask motifs ("-mask <motif file>")
Optimize motifs ("-opt <motif file>")
Dump FASTA files ("-dumpFasta")
Actually, this usually
doesn't
matter that
much. Since
HOMER is a differential motif discovery algorithm,
common repeats are
usually in both the target and background
sequences. However, it
is not uncommon that a transcription factor binds to a
certain class of
repeats, which may cause several large stretches of
similar sequence to
be processed, biasing the results. Usually it's
safer to go with
the masked version.
Region Size ("-size <#>", "-size <#>,<#>", "-size given", default: 200)
The size of the region
used for
motif finding is important. If analyzing ChIP-Seq
peaks from a
transcription factor, Chuck would recommend 50 bp for
establishing the
primary motif
bound by a given transcription factor and 200 bp for
finding both
primary and "co-enriched" motifs for a transcription
factor. When
looking at histone marked regions, 500-1000 bp
is probably a good idea (i.e. H3K4me or H3/H4 acetylated
regions). In theory, HOMER can work with very
large regions (i.e.
10kb), but with the larger the regions comes more
sequence and longer
execution time. These regions will be based off
the center of the peaks. If you prefer an offset,
you can specify "-size -300,100" to search a region of
size 400 that is centered 100 bp upstream of the peak
center (useful if doing motif finding on putative TSS
regions). If you have variable length regions, use
the option "-size given" and HOMER will use the exact
regions that were used as input.
Motif length ("-len <#>" or "-len <#>,<#>,...", default 8,10,12)
Specifies
the
length
of
motifs
to
be
found.
HOMER
will find motifs of each size
separately and then combine the results at the
end. The length of
time it takes to find motifs increases greatly with
increasing
size. In general, it's best to try out enrichment
with shorter
lengths (i.e. less than 15) before trying longer
lengths. Much
longer motifs can be found with HOMER, but it's best to
use smaller sets of sequence when trying to find long
motifs (i.e. use
"-len 20 -size 50"), otherwise it may take way too long
(or take too
much memory). The other trick to reduce the total
resource
consumption is to reduce the number of background
sequences (-N
<#>).
Mismatches allowed in global optimization phase ("-mis <#>", default: 2)
HOMER looks for promising
candidates by initially checking ordinary oligos for
enrichment,
allowing mismatches. The more mismatches you
allow, the more
sensitive the algorithm, particularly for longer
motifs.
However, this also slows down the algorithm a bit.
If searching
for motifs longer than 12-15 bp, it's best to increase
this value to at
least 3 or even 4.
Number of CPUs to use ("-p <#>", default 1)
HOMER is now multicore
compliant. It's not perfectly parallelized,
however, certain
types of analysis can benefit. In general, the
longer the length
of the motif, the better the speed-up you'll see.
Number of motifs to find ("-S <#>", default 25)
Specifies
the
number
of
motifs
of
each
length
to
find. 25 is already quite a bit. If
anything, I'd recommend
reducing this number, particularly for long motifs to
reduce the total
execution time.
Normalize CpG% content instead of GC% content ("-cpg")
Consider tying if HOMER is
stuck
finding "CGCGCGCG"-like motifs. You can also play
around with
disabling GC/CpG
normalization ("-noweight").
Region level autonormalization ("-nlen <#>", default 3, "-nlen 0" to disable)
Motif level autonormalization (-olen <#>, default 0 i.e. disabled)
Autonormalization attempts
to
remove sequence bias from lower order oligos (1-mers,
2-mers ... up to
<#>). Region level autonormalization, which
is for 1/2/3
mers by default, attempts to normalize background
regions by adjusting
their weights. If this isn't getting the job done
(autonormalization is not guaranteed to remove all
sequence bias), you
can try the more aggressive motif level
autonormalization (-olen
<#>). This
performs the autonormalization routine on the oligo
table during de
novo motif discovery. (see here
for more info)
User defined background regions ("-bg <peak file of background regions>")
Why let HOMER randomly
pick you
background regions when you can choose them yourself!!
These will still
be
normalized for CpG% or GC% content just like randomly
chosen sequences
and autonormalized unless these options are turned off
(i.e. "-nlen 0
-noweight"). This can be very useful since HOMER
is a
differential motif discovery algorithm. For
example, you can give
HOMER a set of peaks co-bound by another factor and
compare them to the
rest of the peaks. HOMER will automatically check
if the
background peaks overlap with the target peaks using mergePeaks, and
discard overlapping
regions.
Hypergeometric enrichment scoring ("-h")
By default, findMotifsGenome.pl
uses the binomial distribution to score motifs.
This works well
when the number of background sequences greatly out
number the target
sequences - however, if you are using "-bg"
option
above,
and
the
number
of
background
sequences
is
smaller
than
target sequences, it is a good idea to use the
hypergeometric
distribution instead ("-h").
FYI
-
The
binomial
is
faster
to compute, hence it's use for motif
finding in large numbers of regions.
Find enrichment of individual oligos ("-oligo").
This
creates
output
files
in
the
output
directory
named
oligo.length.txt.
Force findMotifsGenome.pl to re-preparse genome for the given region size ("-preparse").
In case there is a problem
with
the existing preparsed files, force them to be remade
with "-preparse".
Only search for motifs on + strand ("-norevopp")
By default, HOMER looks
for
transcription factor-like motifs on both strands.
This will force
it to only look at the + strand (relative to the peak,
so - strand if
the peak is on the - strand).
Search for RNA motifs ("-rna")
If looking at RNA data
(i.e.
Clip-Seq or similar), this option will restrict HOMER to
only search
the + strand (relative to the peak), and will output RNA
motif logos
(i.e. U instead of T). It will also try to compare
found motifs
to an RNA motif database, which sadly, only contains
miRNAs right
now...
I guess chuck roundhouse kicked all of the splicing and
other RNA
motifs into hard to find databases.
Mask motifs ("-mask <motif file>")
Mask the motif(s) in the
supplied
motif file before starting motif finding. Multiple
motifs can be
in the motif file.
Optimize motifs ("-opt <motif file>")
Instead of looking for
novel de
novo motifs, HOMER will instead try to optimize the
motif
supplied. This is cool when trying to change the
length of a
motif, or find a very long version of a given
motif. For example,
if you specify "-opt <file>" and "-len 50", it
will try to expand
the motif to 50bp and optimize it.
Dump FASTA files ("-dumpFasta")
Like the fact that HOMER
organizes and extracts your sequence files, but don't
care for HOMER as
a motif finding algorithm? That's cool, just
specify "-dumpFasta"
and the files "target.fa" and "background.fa" will show
up in your
output directory. You can then use them with MEME
or
whatever. Just remember, Chuck knows where you
live...
Finding Instance of Specific Motifs
By default, HOMER does not
return
the locations of each motif found in the motif discovery
process.
To recover the motif locations, you must first select the
motifs you're
interested in by getting the "motif file" output by
HOMER. You
can combine multiple motifs in single file if you like to
form a "motif
library". To identify motif locations, you have two
options:
1. Run findMotifsGenome.pl
with the "-find
<motif file>"
option. This will output a tab-delimited text file
with each line
containing an instance of the motif in the target
peaks. The
output is sent to stdout.
For example: findMotifsGenome.pl
ERalpha.peaks hg18
MotifOutputDirectory/ -find motif1.motif >
outputfile.txt
The output file will contain the columns:
2. Run annotatePeaks.pl
with
the "-m <motif
file>"
option (see the annotation
section for
more info). Chuck prefers doing it this way.
This will
output a tab-delimited text file with each line
containing a
peak/region and a column containing instance of each
motif separated by
commas to stdoutThe output file will contain the columns:
- Peak/Region ID
- Offset from the center of the region
- Sequence of the site
- Name of the Motif
- Strand
- Motif Score (log odds score of the motif matrix,
higher
scores are better matches)
For example: annotatePeaks.pl
ERalpha.peaks hg18 -m
motif1.motif > outputfile.txt
The output file will contain columns:
19. CpG%
20. GC%
21. Motif Instances
...
Motif Instances have the following format:
The output file will contain columns:
- Peak/Region ID
- Chromosome
- Start
- End
- Strand of Peaks
19. CpG%
20. GC%
21. Motif Instances
...
Motif Instances have the following format:
<distance from
center of
region>(<sequence>,<strand>,<conservation>)
i.e -29(TAAATCAACA,+,0.00)
i.e -29(TAAATCAACA,+,0.00)
You can also find histogram
of
motif density this way by adding "-hist <#>" to the
command. For example:
Graphing the output with EXCEL:
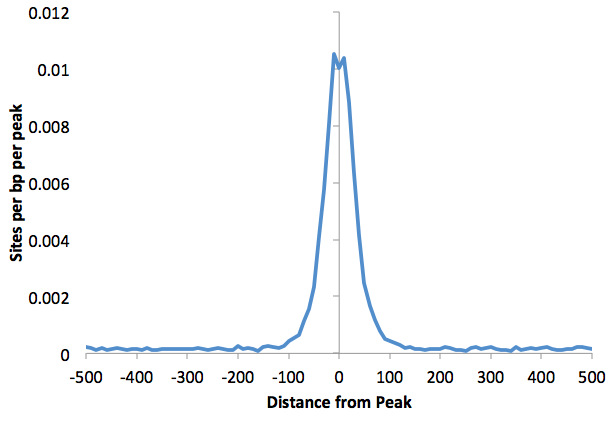
annotatePeaks.pl
ERalpha.peaks
hg18 -m ere.motif foxa1.motif -size 1000 -hist 10
> outputfile.txt
Graphing the output with EXCEL:

Command-line options for findMotifsGenome.pl
Program will find de novo and known motifs in regions in the genomeUsage: findMotifsGenome.pl <pos file> <genome> <output directory> [additional options]
Example: findMotifsGenome.pl peaks.txt mm8r peakAnalysis -size 200 -len 8
Possible Genomes:
...
Custom: provide the path to genome FASTA files (directory or single file)
Heads up: will create the directory "preparsed/" in same location.
Basic options:
-bg <background position file> (genomic positions to be used as background, default=automatic)
removes background positions overlapping with target positions
-chopify (chop up large background regions to the avg size of target regions)
-len <#>[,<#>,<#>...] (motif length, default=8,10,12) [NOTE: values greater 12 may cause the program
to run out of memory - in these cases decrease the number of sequences analyzed (-N),
or try analyzing shorter sequence regions (i.e. -size 100)]
-size <#> (fragment size to use for motif finding, default=200)
-size <#,#> (i.e. -size -100,50 will get sequences from -100 to +50 relative from center)
-size given (uses the exact regions you give it)
-S <#> (Number of motifs to optimize, default: 25)
-mis <#> (global optimization: searches for strings with # mismatches, default: 2)
-norevopp (don't search reverse strand for motifs)
-nomotif (don't search for de novo motif enrichment)
-rna (output RNA motif logos and compare to RNA motif database, automatically sets -norevopp)
Scanning sequence for motifs
-find <motif file> (This will cause the program to only scan for motifs)
Known Motif Options/Visualization
-bits (scale sequence logos by information content, default: doesn't scale)
-nocheck (don't search for de novo vs. known motif similarity)
-mcheck <motif file> (known motifs to check against de novo motifs,
default: /bioinformatics/homer/data/knownTFs/all.motifs
-float (allow adjustment of the degeneracy threshold for known motifs to improve p-value[dangerous])
-noknown (don't search for known motif enrichment, default: -known)
-mknown <motif file> (known motifs to check for enrichment,
default: /bioinformatics/homer/data/knownTFs/known.motifs
Sequence normalization options:
-gc (use GC% for sequence content normalization, now the default)
-cpg (use CpG% instead of GC% for sequence content normalization)
-noweight (no CG correction)
Advanced options:
-h (use hypergeometric for p-values, binomial is default)
-N <#> (Number of sequences to use for motif finding, default=max(50k, 2x input)
-noforce (will attempt to reuse sequence files etc. that are already in output directory)
-local <#> (use local background, # of equal size regions around peaks to use i.e. 2)
-redundant <#> (Remove redundant sequences matching greater than # percent, i.e. -redundant 0.5)
-mask <motif file1> [motif file 2]... (motifs to mask before motif finding)
-opt <motif file1> [motif file 2]... (motifs to optimize or change length of)
-refine <motif file1> (motif to optimize)
-rand (randomize target and background sequences labels)
-ref <peak file> (use file for target and background - first argument is list of peak ids for targets)
-oligo (perform analysis of individual oligo enrichment)
-dumpFasta (Dump fasta files for target and background sequences for use with other programs)
-preparse (force new background files to be created)
-keepFiles (keep temporary files)
homer2 specific options:
-homer2 (use homer2 instead of original homer, default)
-nlen <#> (length of lower-order oligos to normalize in background, default: -nlen 3)
-nmax <#> (Max normalization iterations, default: 160)
-olen <#> (lower-order oligo normalization for oligo table, use if -nlen isn't working well)
-p <#> (Number of processors to use, default: 1)
-e <#> (Maximum expected motif instance per bp in random sequence, default: 0.01)
-cache <#> (size in MB for statistics cache, default: 500)
-quickMask (skip full masking after finding motifs, similar to original homer)
Original homer specific options:
-homer1 (to force the use of the original homer)
-depth [low|med|high|allnight] (time spent on local optimization default: med)

Can't figure something out? Questions, comments, concerns, or other feedback:
cbenner@salk.edu